A novel mutation of CAPN1 in patients with complicated hereditary spastic paraplegia
Lilu Kuang1,†, Liang Deng1,†, Jianhuang Chen1,†, Runcheng He2*
¹Department of Neurology, Jili Hospital, Changsha, Hunan 410399, China.
²Department of Neurology, the Second Xiangya Hospital, Central South University, Changsha, Hunan 410011, China.
†These authors have contributed equally to this work and are co-first authors.
*Corresponding author
*Runcheng He. Department of Neurology, the Second Xiangya Hospital, Central South University, Changsha, Hunan, P.R. China, 410011. E-mail: heruncheng_xy@163.com
DOI: 10.55920/JCRMHS.2024.07.001305
Figure 1: Brain Magnetic resonance imaging (MRI) showed the atrophy of the cerebellum, and spinal cord MRI showed thoracic cord atrophy
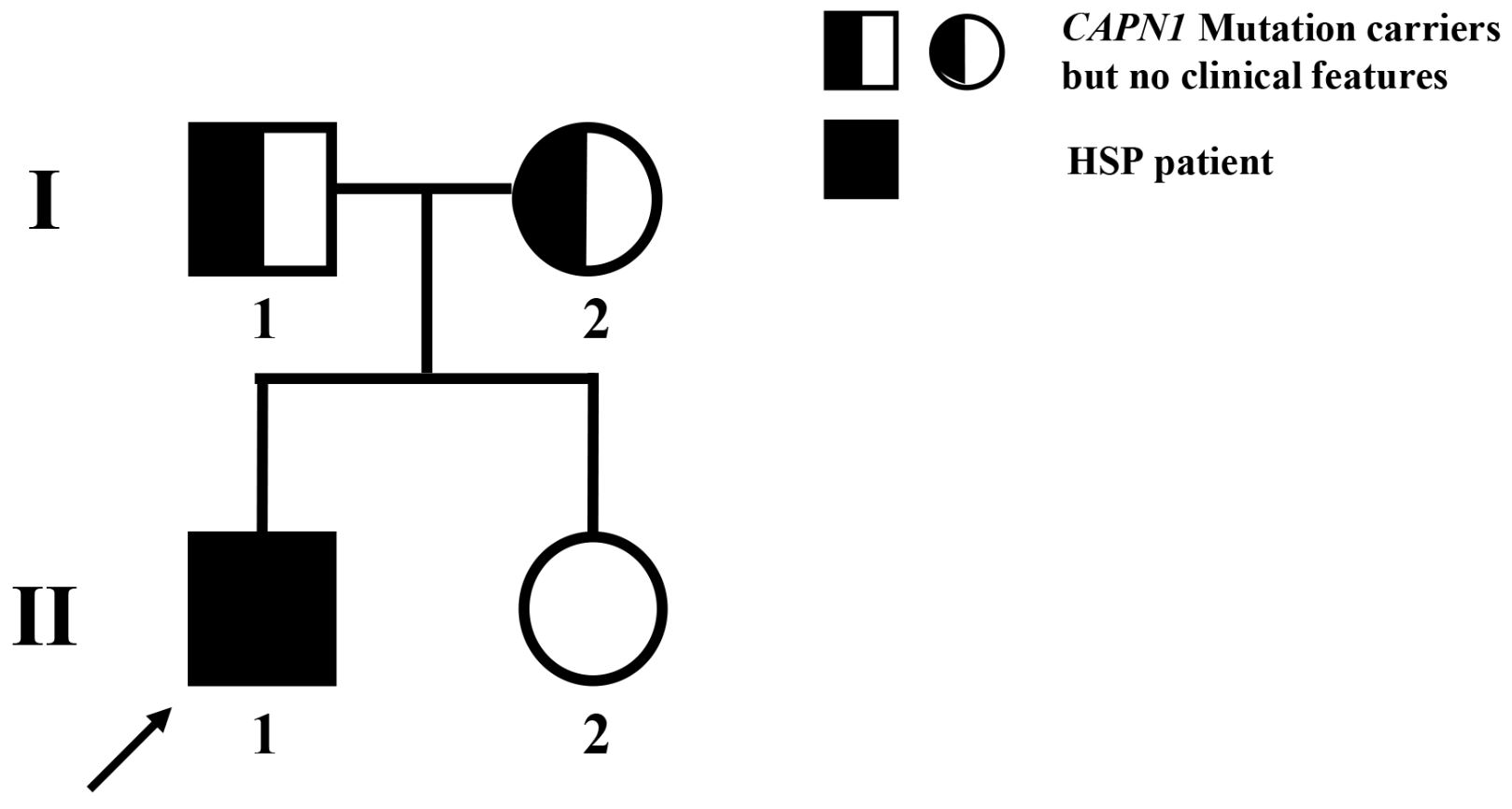
Figure 2: Pedigree of the family. Patient I-1 is the proband.
His sister and parents were asymptomatic (Figure 2). Genomic DNA was extracted from peripheral blood of both patients and his parents or sister through a standard phenol-chloroform method. All family members signed a written informed consent. Multiple molecular genetic tests of the proband were used, including long-range polymerase chain reaction (PCR), target sequencing, and Sanger sequencing. A total of 10 fragments including CAG/CTG expansion sequences were amplified by PCR in different reactions using an ABI Prism Thermal cycler. At the same time, the proband was performed by target sequencing of a custom panel containing over 730 virulence genes known to cause HSP, SCA and other movement disorders. The sequencing was carried out by Illumina HiSeq X-10 platform. The mean depth of the targeted regions was 161.43X and the coverage was 99.88%. Subsequently, we performed Sanger sequencing to confirm the candidate variants detected through targeted sequencing analysis. These variants were also performed in his parents and his sister to confirm the family co-segregation.
Figure 3: DNA sequence at codon 480 of CAPN1 gene from the proband (A), I-1(B), I-2(C),and II-2 (D). The red arrow indicates the c.1439T>C mutation causing a substitution of Leucine by Proline at codon 480 of the protein.
The mutation is predicted to be “deleterious” by sorts intolerant from tolerant (SIFT), and “probably damaging” by Polyphen-2. The variant meets the criteria of PS2, PM2, and PP3 of the American College of Medical Genetics (ACMG) guideline, and its pathogenicity is further supported.
Therefore, we conclude that this missense variant is a “likely pathogenic” variant for SPG76.