Mechanism Study of Guishen Decoction in Treating Ischemic Stroke Based on Network Pharmacology and Molecular Docking
Yan Song LIU1,2, Yu Chen LIU1, Zhi Guang SONG1, Wang Hua LIU1,2,3*
¹Provincial Key Laboratory of TCM Diagnostics, Hunan University of Chinese Medicine, Changsha 410208, Hunan, China
²Key Laboratory of TCM Heart and Lung Syndrome Differentiation & Medicated Diet and Dietotherapy, Hunan University of Chinese Medicine, Changsha 410208, Hunan, China
³Hunan Engineering Technology Research Center For Medicinal and Functional Food, Hunan University of Chinese Medicine, Changsha 410208, Hunan, China
*Corresponding author
*Wang-hua LIU, Provincial Key Laboratory of TCM Diagnostics, Hunan University of Chinese Medicine, Changsha 410208, Hunan, China
DOI: 10.55920/JCRMHS.2025.10.001425
Figure 1: Network pharmacology and molecular docking workflow of bioactive compounds in Guishen Decoction for the treatment of ischemic stroke.
Figure 2: The target genes of ischemic stroke based on Genecards, Online Mendelian Inheritance in Man (OMIM), The Pharmacogenomics Knowledgebase (PharmGkb), Therapeutic Target Database (TTD) and DrugBank databases.
Figure 3: The potential targets of between bioactive compounds of Guishen Decoction against ischemic stroke via Venn diagram. There are 168 target genes in Guishen Decoction bioactive compounds and ischemic intersection (the middle part).
Construction of Venn Diagram
Use the venn package of R 4.3.2 software to examine the targets of the ingredients in Guishen Decoction and the targets associated with ischemic stroke, and clarify the drug ingredient targets of Guishen Decoction that are related to ischemic stroke, the findings of the analysis are illustrated using a Venn diagram.
Construction of “Compound-Target-Disease” Network
Use Cytoscape 3.10.1 (https://cytoscape.org/, accessed on 23 February 2024) software to illustrate the interaction between the drug ingredients of Guishen Decoction and ischemic stroke. The specific method is: classify and adjust the activated ingredient information of the compositions in Guishen Decoction and integrate the data into Cytoscape software enables the active ingredients of drugs to be visually displayed in Cytoscape; taking the drug ingredients of Guishen Decoction and the common target genes of ischemic stroke as network nodes, the relationship between the active ingredients of the drug and the common target genes is Displayed in the form of connecting lines, a “compound-target-disease” network is formed.
Protein-Protein-Interaction (PPI) Network Analysis
The STRING 12.0 (https://cn.string-db.org/, accessed on 23 February 2024) database was used to analyze the intersection of bioactive compounds of Guishen Decoction and ischemic stroke target genes, and the analysis content was limited to human species. , the confidence level is 0.7 and above, perform network visualization on the obtained protein, and enter the network into Cytoscape software. After integrating the data from the STRING database on the Cytoscape platform, use the CytoHubba plug-in (Chin et al., 2014) to calculate the maximum clique centrality (MCC), maximum neighborhood component (MNC) and degree values of each target gene, respectively determine the highest-ranking 10 genes with three algorithm scores, thereby determining potential central target genes in the network. The results of these three algorithms were copied into a set of target genes that were used as potential key targets in the network.
Gene Ontology and Pathway Enrichment Analysis
For the purpose of exploring the potential mechanism of action between the medicinal ingredients of Guishen Decoction and ischemic stroke, the Database for Annotation, Visualization, and Integrated Discovery database (DAVID), version 2021 (https://david.ncifcrf.gov/home.jsp, accessed on 25 February 2024) (Sherman et al., 2022) was deployed to analyze biological functions, cellular structures, molecular activities and signaling cascades (Dennis et al., 2003; Huang et al., 2009; Kanehisa et al., 2023) to obtain GO, and the first 15 results of KEGG analysis, using the Bioinformatic (https://www.bioinformatics.com.cn, accessed on 25 February 2024) platform to visualize the results. After calculating the P value, statistically significant results have a P value of less than 0.05.
Molecular Docking
To analyse the interaction between the Guishen Decoction and the main genes specified via the UniProt database molecular connection is used to find the UniProt ID of the main gene., and the crystal structure of the core gene was searched in the RCSB protein database (https://www.rcsb.org/, accessed on 25 February 2024) (Berman et al., 2000) based on the protein ID, resulting in IL6 (UniProt ID: P05231; PDB ID: 1ALU), TNF (UniPort ID: P01375; PDB ID: 1TNF), IL1B (UniProt ID: P01584; PDB ID: 1HIB). Import the three-dimensional structure of the core gene into PyMOL 2.5 (https://pymol.org/, accessed on 25 February 2024) software, remove water molecules and existing small molecule ligands, and obtain the protein ligand file; use the PubChem database (https://pubchem.ncbi.nlm.nih.gov/, accessed on 25 February 2024) (Kim et al., 2021) to search Guishen Decoction active ingredients matrine, quercetin, kaempferol For 2D structure, ChemOffice 22 (https://www.chemdraw.com.cn/, accessed on 25 February 2024) software was used to convert the 2D structure of the active ingredient into a three-dimensional structure, and the three-dimensional structure was optimized using minimum energy. Using the active ingredient as the ligand and the core gene as the receptor, AutoDockTools 4.2 (http://autodock.scripps.edu/, accessed on 25 February 2024) (Morris et al., 2009) software was used to predict the binding conformation between the ligand and the receptor, and AutoDockVina 1.2.0 (https://vina.scripps.edu/, accessed on 25 February 2024) (Eberhardt et al., 2021; Trott & Olson, 2010) software was used to visualize the molecular interaction between the protein and the ligand.
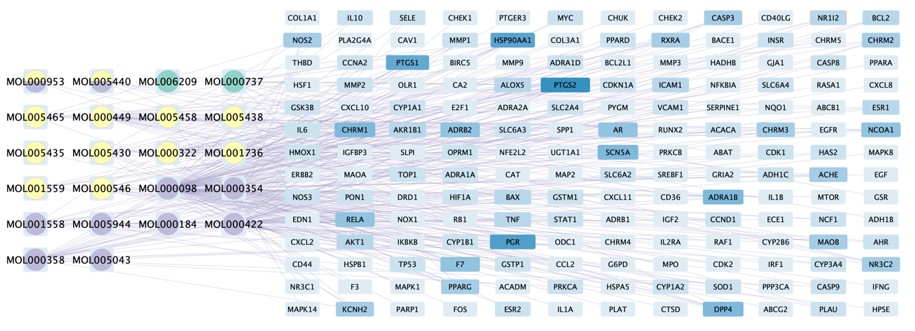
Figure 4: The compound-target network of Guishen Decoction bioactive compounds against ischemic stroke by using Cytoscape. The blue nodes represent the interacting target genes between the compound (yellow, green and purple nodes) and the ischemic stroke.
Table 1: The pharmacokinetic properties of the bioactive ingredients of Guishen Decoction based on Traditional Chinese Medicine Systems Pharmacology (TCMSP) database and Uniprot database.
Gene Ontology and Pathway Enrichment Analysis
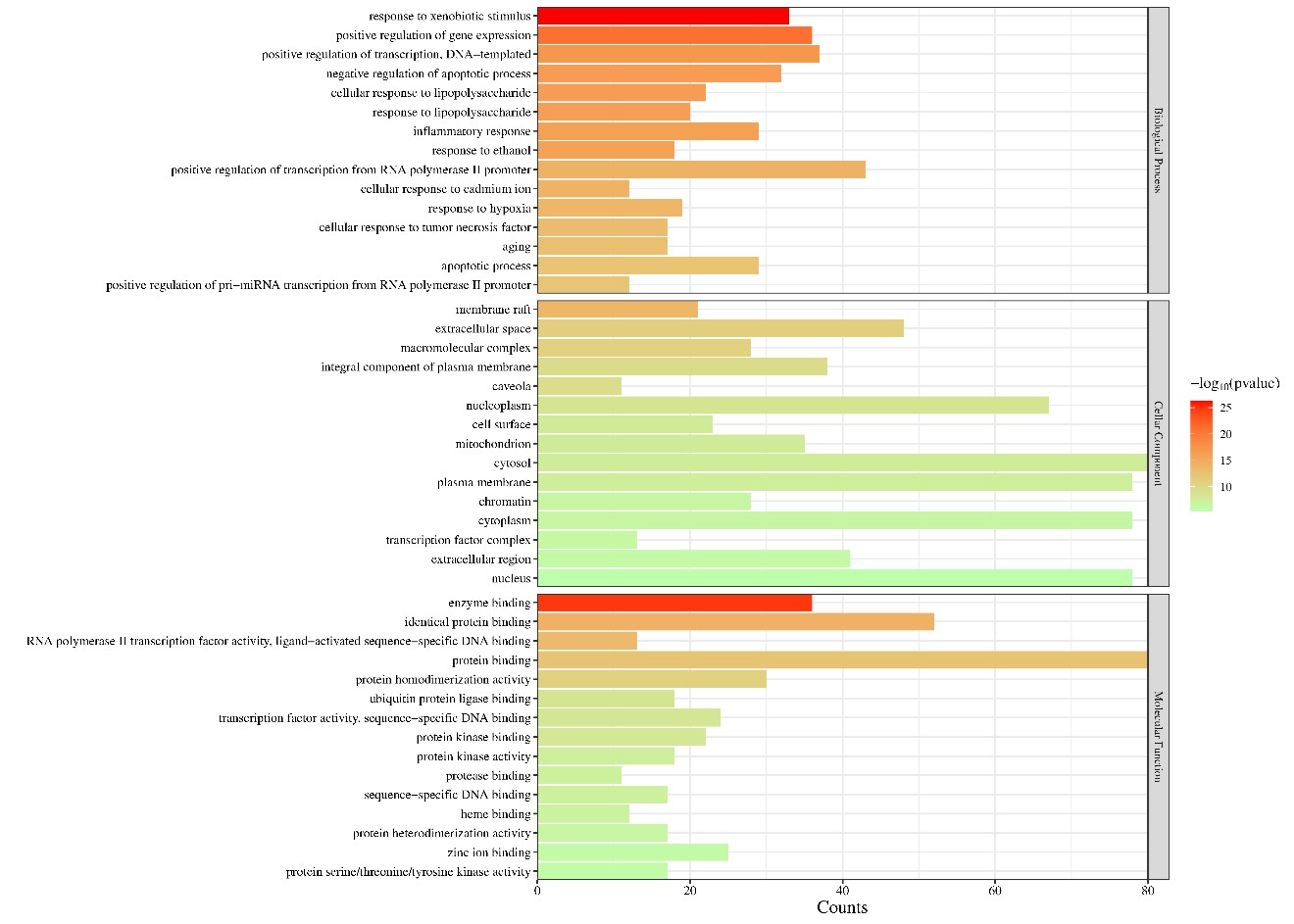
In order to further understand the biological processes, cellular components, molecular functions and pathways involved in the therapeutic effect of Guishen Decoction bioactive compounds on ischemic stroke, a Gene Ontology (GO) study and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis was conducted using DAVID bioinformatics resources and the Bioinformatics platform. The results of GO analysis (Figure 7) show that the targets of the bioactive compounds in Guishen Decoction are the most significant biological processes related to xenobiotic stimulus and lipopolysaccharide. Analysis of cellular components showed that these targets mainly involved membrane raft, membrane microdomain and caveola. In terms of molecular function, the top 10 significantly enriched terms include DNA-binding transcription factor binding, G protein-coupled amine receptor activity and transcription coregulator binding. The results of KEGG analysis (Figure 8) show that many targets of the bioactive compounds of Guishen Decoction on ischemic stroke are enriched in signaling pathways, such as lipid and atherosclerosis, AGE-RAGE signaling pathway in diabetic complications, fluid shear stress and atherosclerosis and prostate cancer. Through the analysis of metabolic pathways, signaling pathways such as pathways of neurodegeneration are directly related to ischemic stroke. Find the potential targets and mechanisms of Guishen Decoction to treat ischemic stroke by regulating Pathways of neurodegeneration (Figure 9).
Figure 5: Protein-protein interaction networks are constructed using the STRING database. The network nodes represent proteins, each node represents all the proteins produced by a single, protein-coding gene locus. The colored nodes represent query proteins and first shell of interactors, and the white nodes representing the second shell of interactors. The edges represent protein-protein associations, the confidence levels of interaction are visually represented in the network diagram using color edges, including from curated databases, experimentally determined, gene neighborhood, gene fusions, gene co-occurrence, textmining, co-expression and protein homology.
Figure 6: By combining 3 algorithms, MCC, MNC, degree, the key gene network of Guishen Decoction bioactive compounds with scores above the median for ischemic stroke was screened out. The genes with the highest values are considered the most important key genes and are indicated in yellow. Conversely, genes with lower values are considered less important and are indicated in blue, indicating each gene’s ranking position in the network.
Molecular Docking
Through molecular docking research, the interaction relationship between the three active compounds selected in Guishen Decoction (matrine, kaempferol, quercetin) and three potential target genes (IL6, IL1B, TNF) was analyzed. AutoDockTools software was used to conduct molecular docking studies, and a total of 9 docking results were obtained, 7 of which had binding energies that resisted -5.0 kcal/mol. The docking scores are shown in Table 2. The lower the binding energy, the tighter the binding between the compound and the protein. The binding energy of kaempferol and quercetin to each target gene is lower than -5.0 kcal/mol. Therefore, kaempferol and quercetin are important in the treatment of ischemic stroke. Play a particularly important role. The results show that most active ingredients have strong binding ability to the target gene TNF. The docking scores of kaempferol, matrine and quercetin combined with TNF are -6.6, -4.9 and -6.7 respectively. This result suggests that the bioactive compounds target TNF of Guishen Decoction may play a key role in the treatment of ischemic stroke. Use PyMOL to construct visualizations of the lowest binding energies between targets and bioactive compounds.
Figure 7: Gene ontology (GO) enrichment analysis. The bar chart represents the most significantly enriched GO terms in biological processes, cellular components, and molecular function, comprising the top 10 terms related to the target genes.
Figure 8: Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis. The bar chart visualizes the top 30 enriched KEGG pathways of Guishen Decoction against ischemic stroke.
Table 2: Binding energy between active compounds and three core targets of Guishen Decoction
Figure 9: Potential targets and mechanism of bioactive compounds in Guishen Decoction against ischemic stroke. The figure represents the neurotrophin signaling pathway. The red ndoes represent the genes targeted within the pathways of neurodegeneration.
Figure 10: Molecular docking results of the lowest binding energy in each target with the bioactive compound of Guishen Decoction (a) IL6-quercetin, (b) IL1B-quercetin, (c) TNF-quercetin.
After network pharmacology analysis, this study identified 191 potential bioactive compound targets of Guishen Decoction; in the network diagram, this article focused on three central targets: IL6, IL1B, and TNF. To elucidate the pharmacokinetic characteristics of Guishen Decoction bioactive decoder compounds in ischemic brain injury, this study used oral bioavailability (OB) ≥ 30% and drug-likeness (DL) ≥ 0.18 as the screening criteria, and Discard the remaining compounds whose corresponding targets cannot be obtained, and only summarize the remaining compounds as potential bioactive compounds. Three compounds, matrine, quercetin, and kaempferol, were selected from Guishen Decoction for further research because they meet the criteria for drug screening and may play an important role in the treatment of ischemic stroke.
Interpreting the KEGG pathway results in the process of GO and KEGG enrichment analysis shows that Apoptosis, Neurotrophin signaling pathway, Pathways of neurodegeneration, Glioma, lipids, IL-17 signaling pathway, HIF-1 signaling pathway, etc. have all been confirmed and lacking. (Cui et al., 2021; Ghosh et al., 2019; Yang C et al., 2019), this article conducts visual research on related signaling pathways, and conducts molecular docking verification of 3 key targets and 3 active ingredients. The results show that quercetin and kaempferol play an important role in the treatment of ischemic stroke, as these compounds can be closely bound in the active pouch of key targets with low binding energy. At the same time, TNF looks relatively well with most of guishen's active analgesics.
This article uses network pharmacology and molecular docking to study the potential active ingredients, possible relevant targets and important biological pathways of Guishen Decoction in the treatment of ischemic stroke, providing a preliminary theory for further experimental research. It must be noted that the pharmacological mechanisms proposed in this article are all derived from network pharmacology technology and require further verification by pharmacological and clinical studies.